Discovery & Development: Genomics
Designing genetic control: synthetic promoters powering modern therapeutics and biomanufacturing
How is the synthetic design of genetic elements, such as promoters for cell and gene therapies and biologics, improving development and manufacturing?
Priyanka Anujan at SynGenSys
Across biological systems, the unravelling of the complexities of gene expression regulation has evolved from being an interesting experimental pursuit, to being an engineering foundation for next-generation biomanufacturing and therapeutic outcomes. Initial genetic engineering techniques predominantly relied on gene overexpression or knockout strategies to demonstrate functional outcomes. Although these methods sufficed for proof-of-concept studies, they often imposed metabolic burdens, destabilised endogenous regulatory networks and diminished cellular fitness.1,2
As the fields of synthetic biology, metabolic engineering and cell and gene therapy (CGT) have progressed, it has become increasingly evident that cellular systems are inherently dynamic, resource-limited and highly interconnected. Excessive, insufficient or poorly timed gene expression can disrupt pathway balance, reduce productivity and, in therapeutic applications, compromise safety.
Synthetic, regulatable genetic elements mitigate this limitation by providing programmable control over the timing, location and intensity of gene expression.3 These include engineered promoters and enhancers, modular transcription factor architectures, synthetic response elements and chromatin-targeting regulators that facilitate graded, inducible or tissue-specific transcriptional activity.4,5 Rather than viewing gene expression as a binary switch, contemporary design strategies enable the modulation of expression levels over extensive dynamic ranges while ensuring stability and reproducibility.
The urgency surrounding precise gene regulation reflects the increasing complexity of applications. In industrial biomanufacturing, the coordinated modulation of multiple genes is crucial for balancing cellular growth with product yield.2 In synthetic biology, the development of gene circuits – such as logic gates, oscillators and feedback controllers – relies on predictable transcriptional components with controllable variability.5 In therapeutic applications, programmable gene regulation platforms facilitate the selective modulation of disease-relevant genes with adjustable intensity, thereby enhancing safety margins compared to constitutive viral promoters.3,6 Advances in massively parallel reporter assays (MPRAs) and machine learning (ML) have further expanded the design space, enabling the systematic characterisation of regulatory DNA and its de novo generation with defined transcriptional outputs.7,8
Together, these developments have transformed gene expression from a coarse on/off mechanism into a programmable engineering layer that underpins next-generation biomanufacturing and precision medicine (Figure 1).
From natural constraints to engineered control
Traditional promoters, whether endogenous or viral, have historically served as foundational elements in genetic engineering. Nevertheless, as the scope of applications necessitates heightened precision, the constraints of these promoters have become progressively apparent.
A persistent concern is unintended basal or off-target activity. Widely used mammalian promoters, including cytomegalovirus (CMV) and simian virus 40 promoters and their derivatives, drive strong constitutive expression but offer limited spatial or cell-type specificity.4 In therapeutic settings, broad activity can reduce the on-target/off-target expression ratio, narrowing safety margins. Viral promoters are also susceptible to epigenetic silencing and methylation-dependent inactivation, leading to variability or loss of expression over time.6 Although tissue-specific promoters improve spatial restriction, they often lack sufficient strength for high-level therapeutic output, and may still exhibit residual expression in non-target cells.6
Dynamic range and tunability present additional constraints. In yeast and mammalian systems, native promoters frequently do not encompass the extensive transcriptional gradients necessary to equilibrate multi-gene metabolic pathways.1,2 Minor architectural modifications can induce disproportionate changes in activity, thereby complicating predictive design.5 Furthermore, promoter output is influenced by factors such as genomic integration site, copy number, surrounding regulatory elements and cellular state, which constrain their portability across different platforms.1
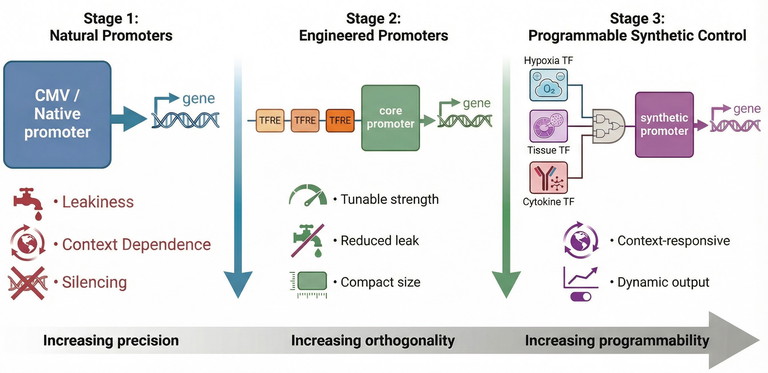
Figure 1: Reprogramming gene expression: from natural promoters to synthetic regulatory systems
In adeno-associated virus (AAV) gene therapy, packaging constraints exacerbate these challenges. With an effective cargo capacity of approximately 4.7kb, large natural promoters considerably limit the available space for therapeutic genes and regulatory safeguards.6 Truncating sequences to meet size restrictions can undermine their efficacy or specificity, frequently necessitating reliance on compact viral promoters that may be leaky or susceptible to silencing.
Finally, native promoters are integrated within endogenous transcriptional networks, which restrict orthogonality and elevate the likelihood of transcription factor competition or regulatory crosstalk when expressing multiple transgenes.5,9 As therapeutic constructs grow in complexity, these limitations hinder scalability and reproducibility.
Collectively, these illustrate why traditional promoters, although valuable workhorses, often fall short in meeting the demands of contemporary high-precision synthetic biology and translational applications.
The rise of synthetic promoters
Synthetic promoters have emerged as engineered solutions to overcome the limitations of native regulatory elements.
Rather than relying on naturally occurring sequences, they are built by deliberately rearranging and designing cis-regulatory elements (CREs) and core promoter architectures to achieve defined strength, inducibility and specificity.4,5 Combinatorial assembly of transcription factor regulatory elements (TFREs), optimisation of element spacing and integration with minimal core promoters enable precise control of transcriptional output. In mammalian systems, MPRA-based platforms have enabled screening of thousands of compact synthetic promoters, spanning dynamic ranges of 50-100-fold in some contexts.7 Large synthetic promoter libraries have further demonstrated multilevel regulation within multi-gene constructs while minimising cross-interference between expression cassettes.8
Host- and cell-type-specific design strategies have further accelerated adoption. In Chinese hamster ovary (CHO) cells, TFRE-guided synthetic promoter libraries achieved up to two-fold higher activity than CMV and provided greater than 100-fold tunability.9,10 Controlled modulation of heavy and light chain expression has been shown to improve recombinant protein quality and reduce aberrant assembly in complex biologics.10
In gene therapy, compact synthetic promoters engineered for retinal pigment epithelial cells demonstrated activity surpassing that of endogenous BEST1 and, in certain configurations, approaching the levels of CMV, while maintaining specificity.11 Multi-responsive promoters, which incorporate hypoxia- and inflammation-sensitive elements, have exhibited additive or synergistic activation, thereby facilitating disease-context-dependent gene expression.12

In the field of cellular immunotherapy, transcription factor-responsive synthetic promoters have been engineered to identify activation and exhaustion states in primary human T-cells, facilitating the context-dependent regulation of therapeutic genes.13 Promoters that respond to the tumour microenvironment further limit effector gene expression to locations of inflammation or hypoxia, thereby improving safety profiles in CAR T-cell therapies.12,14
These advances exemplify a broader transition from utilising naturally occurring regulatory elements to designing bespoke transcriptional modules that can be quantitatively calibrated, assembled and incorporated into progressively more complex genetic circuits.
Applications across therapeutic areas
Biopharmaceutical manufacturing
In the production of CHO-based biologics, synthetic promoters facilitate precise regulation of transgene expression, thereby balancing productivity with cellular health.9,10 Tunable promoter libraries permit the optimisation of multi-gene pathways and enhance control over complex protein assembly. Beyond increasing yield, promoter engineering has also demonstrated improvements in product quality attributes, highlighting the economic significance of transcriptional regulation.10
Gene therapy
In AAV-based gene therapy, compact synthetic promoters effectively address the trade-offs between expression strength and specificity, as well as packaging limitations.6 Tissue-specific synthetic promoters designed for applications in the liver, muscle and retina improve targeting precision while conserving vector space.11,15 Tumour-specific and inflammation-responsive promoters confine gene expression to malignant tissues, thereby reducing systemic toxicity.14,16
Cell therapy
In the field of engineered immune cell therapies, synthetic promoters are progressively employed to link therapeutic activity to environmental signals. Multi-input promoters responsive to cytokines, hypoxia or transcriptional state alterations facilitate dynamic regulation of proliferation and effector functions.12,14 These context-aware systems represent a vital advancement towards programmable, state-sensitive CAR T-cell and natural killer cell therapies.13
Artificial intelligence-driven regulatory design
ML and deep generative models trained on MPRA data sets are beginning to predict promoter activity directly from sequence.7,17 Generative adversarial networks and language-model approaches have demonstrated the capacity to design synthetic regulatory DNA with specific transcriptional profiles, thereby broadening the accessible regulatory landscape beyond naturally occurring motifs.17,18
Across manufacturing and therapeutic domains, synthetic promoters function not merely as stronger alternatives to viral elements but as programmable control layers that enable precision design.
Future outlook
The next stage of promoter engineering will be characterised by the integration of computational modelling, high-throughput functional screening and translational validation. As data sets that correlate regulatory sequences with transcriptional output expand, the accuracy and generalisability of predictive models are anticipated to improve.7,17 Incorporating chromatin context, epigenetic features and three-dimensional genome architecture into design frameworks may further augment predictive capabilities.19
In therapeutic contexts, compact multi-input promoters capable of integrating inflammatory, metabolic and hypoxic signals may facilitate autonomous gene circuits that adapt dynamically to cellular states.12,14 Such systems could diminish vector dose requirements, mitigate off-target toxicity and broaden therapeutic windows. Standardisation of benchmarking frameworks, enhanced orthogonality and long-term stability in vivo remain imperative challenges.6,9 Nonetheless, the trajectory is clear: regulatory DNA is progressing from a passive sequence to an actively engineered control system.
Synthetic promoters signify a shift from utilising naturally occurring regulatory components to the development of programmable genetic control systems. As engineering principles integrate with biological insights, gene expression is increasingly regarded as a quantitative, adjustable and context-sensitive parameter, transforming the methodologies employed in constructing cellular factories, engineering immune cells and delivering gene therapies.
References:
1. Blazeck J et al (2012), ‘Promoter engineering: recent advances in controlling transcription at the most fundamental level’, Biotechnology Journal, 7(3), 346-364
2. Tang W et al (2020), ‘Engineering transcriptional control for metabolic pathway optimisation in yeast’, Metabolites, 10(8), 320
3. Ansari A M et al (2025), ‘Programmable gene regulation systems for therapeutic applications’, Journal of Advanced Research, 78, 351-361
4. Cazier A P et al (2021), ‘Advances in promoter engineering: novel applications and predefined transcriptional control’, Biotechnology Journal, 16(8), 2100239
5. Li H S et al (2020), ‘Multidimensional control of therapeutic human cell function with synthetic gene circuits’, Science, 368(6495), eaaz0203
6. Artemyev V et al (2024), ‘Synthetic promoters in gene therapy: design approaches, features and applications’, Cells, 13(19), 1963
7. Zahm A M et al (2024), ‘A massively parallel reporter assay library to screen short synthetic promoters in mammalian cells’, Nature Communications, 15, 10353
8. Fu Z H et al (2025), ‘Synthetic tunable promoters for flexible control of multi-gene expression in mammalian cells’, Journal of Advanced Research, 78, 351-361
9. Brown A J et al (2014), ‘Synthetic promoters for CHO cell engineering’, Biotechnology and Bioengineering, 111(8), 1638-1649
10. Sou S N et al (2023), ‘CHO synthetic promoters improve expression and product quality of biotherapeutic proteins’, Biotechnology Progress, 39(2), e3348
11. Johari Y B et al (2021), ‘Design of synthetic promoters for controlled expression of therapeutic genes in retinal pigment epithelial cells’, Biotechnology and Bioengineering, 118(5), 2001-2015
12. Mohamed H et al (2016), ‘Assembly PCR synthesis of compact multi-responsive promoters suited to gene therapy application’, Scientific Reports, 6, 29388
13. Visit: biorxiv.org/content/10.1101/2025.01.09.63203 4v1
14. Greenshpan Y et al (2021), ‘Synthetic promoters to induce immune effectors into the tumor microenvironment’, Communications Biology, 4, 143
15. Biłas R et al (2016), ‘Cis-regulatory elements used to control gene expression in plants’, Plant Cell Reports, 35, 1059-1071
16. Montaño-Samaniego M et al (2020), ‘Strategies for targeting gene therapy in cancer cells with tumorspecific promoters’, Frontiers in Oncology, 10, 605380
17. Zrimec J et al (2022), ‘Deep learning suggests that gene expression is encoded in all parts of a coevolving interacting gene regulatory structure’, Nature Communications, 13, 3867
18. Karollus A et al (2023), ‘Generic sequence features explain most of the mRNA expression variation in humans’, Genome Biology, 24, 289

Priyanka Anujan PhD is a senior cell engineering scientist at SynGenSys, specialising in functional validation of synthetic regulatory elements for advanced therapeutic and biomanufacturing applications. Her expertise includes cell engineering, viral vector optimisation, and translational evaluation in immune and mammalian cell systems. With a background in molecular biology and cell engineering, she focuses on developing programmable genetic control systems to enhance precision, safety and scalability in next-generation CGTs. Her research integrates experimental biology and synthetic design to create robust, context-responsive therapeutic platforms.